-Search query
-Search result
Showing 1 - 50 of 148 items for (author: wataru & a)

EMDB-38330: 
ETB-eGt complex bound to endothelin-1
Method: single particle / : Oshima HS, Sano FK, Akasaka H, Iwama A, Shihoya W, Nureki O

PDB-8xgr: 
ETB-eGt complex bound to endothelin-1
Method: single particle / : Oshima HS, Sano FK, Akasaka H, Iwama A, Shihoya W, Nureki O

EMDB-38232: 
The cryo-EM structure of the RAD51 L2 loop bound to the linker DNA with the blunt end of the nucleosome
Method: single particle / : Shioi T, Hatazawa S, Ogasawara M, Takizawa Y, Kurumizaka H

PDB-8xbx: 
The cryo-EM structure of the RAD51 L2 loop bound to the linker DNA with the blunt end of the nucleosome
Method: single particle / : Shioi T, Hatazawa S, Ogasawara M, Takizawa Y, Kurumizaka H

EMDB-17740: 
Cryo-EM structure of NR5A2-nucleosome complex SHL+5.5
Method: single particle / : Kobayashi W, Sappler A, Bollschweiler D, Kummecke M, Basquin J, Arslantas E, Ruangroengkulrith S, Hornberger R, Duderstadt K, Tachibana K

EMDB-17741: 
Cryo-EM structure of the nucleosome containing Nr5a2 motif at SHL+5.5
Method: single particle / : Kobayashi W, Sappler A, Bollschweiler D, Kummecke M, Basquin J, Arslantas E, Ruangroengkulrith S, Hornberger R, Duderstadt K, Tachibana K

PDB-8pki: 
Cryo-EM structure of NR5A2-nucleosome complex SHL+5.5
Method: single particle / : Kobayashi W, Sappler A, Bollschweiler D, Kummecke M, Basquin J, Arslantas E, Ruangroengkulrith S, Hornberger R, Duderstadt K, Tachibana K

PDB-8pkj: 
Cryo-EM structure of the nucleosome containing Nr5a2 motif at SHL+5.5
Method: single particle / : Kobayashi W, Sappler A, Bollschweiler D, Kummecke M, Basquin J, Arslantas E, Ruangroengkulrith S, Hornberger R, Duderstadt K, Tachibana K
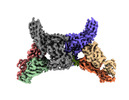
EMDB-34174: 
Structure of beta-arrestin2 in complex with a phosphopeptide corresponding to the human Atypical chemokine receptor 2, ACKR2 (D6R)
Method: single particle / : Maharana J, Sarma P, Yadav MK, Banerjee R, Shukla AK
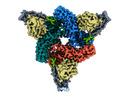
EMDB-36078: 
Structure of beta-arrestin2 in complex with M2Rpp
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK
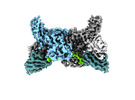
EMDB-36081: 
Structure of beta-arrestin2 in complex with D6Rpp (Local Refine)
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK
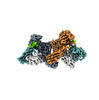
EMDB-36082: 
Structure of beta-arrestin1 in complex with D6Rpp
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK
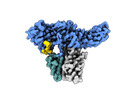
EMDB-36090: 
Structure of Muscarinic receptor (M2R) in complex with beta-arrestin1 (Local refine, cross-linked)
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK
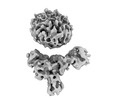
EMDB-36091: 
Muscarinic receptor (M2R) in complex with beta-arrestin1 (Low resolution full map, Cross-linked)
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK
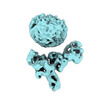
EMDB-36093: 
Muscarinic receptor (M2R) in complex with beta-arrestin1 (Low resolution full map, non-crosslinked)
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK

EMDB-36110: 
Structure of basal beta-arrestin2
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK
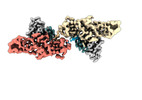
EMDB-36124: 
Structure of beta-arrestin1 in complex with C3aRpp
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK
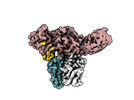
EMDB-36126: 
Structure of Muscarinic receptor (M2R) in complex with beta-arrestin1 (Local Refine, non-cross linked)
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK

PDB-8go9: 
Structure of beta-arrestin2 in complex with a phosphopeptide corresponding to the human Atypical chemokine receptor 2, ACKR2 (D6R)
Method: single particle / : Maharana J, Sarma P, Yadav MK, Banerjee R, Shukla AK

PDB-8j8r: 
Structure of beta-arrestin2 in complex with M2Rpp
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK

PDB-8j8v: 
Structure of beta-arrestin2 in complex with D6Rpp (Local Refine)
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK

PDB-8j8z: 
Structure of beta-arrestin1 in complex with D6Rpp
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK

PDB-8j97: 
Structure of Muscarinic receptor (M2R) in complex with beta-arrestin1 (Local refine, cross-linked)
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK

PDB-8j9k: 
Structure of basal beta-arrestin2
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK

PDB-8ja3: 
Structure of beta-arrestin1 in complex with C3aRpp
Method: single particle / : Maharana J, Sarma P, Yadav MK, Chami M, Banerjee R, Shukla AK

PDB-8jaf: 
Structure of Muscarinic receptor (M2R) in complex with beta-arrestin1 (Local Refine, non-cross linked)
Method: single particle / : Maharana J, Sano FK, Shihoya W, Banerjee R, Nureki O, Shukla AK
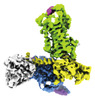
EMDB-35814: 
ETB-Gi complex bound to endothelin-1
Method: single particle / : Sano FK, Akasaka H, Shihoya W, Nureki O

EMDB-35815: 
ETB-Gi complex bound to Endotheline-1, focused on receptor
Method: single particle / : Sano FK, Akasaka H, Shihoya W, Nureki O

PDB-8iy5: 
ETB-Gi complex bound to endothelin-1
Method: single particle / : Sano FK, Akasaka H, Shihoya W, Nureki O

PDB-8iy6: 
ETB-Gi complex bound to Endotheline-1, focused on receptor
Method: single particle / : Sano FK, Akasaka H, Shihoya W, Nureki O

EMDB-32407: 
RNA polymerase II elongation complex bound with Elf1 and Spt4/5, stalled at SHL(-4) of the nucleosome
Method: single particle / : Osumi K, Kujirai T, Ehara H, Sekine S, Takizawa Y, Kurumizaka H

EMDB-32408: 
RNA polymerase II elongation complex bound with Elf1 and Spt4/5, stalled at SHL(-3.5) of the nucleosome
Method: single particle / : Osumi K, Kujirai T, Ehara H, Sekine S, Takizawa Y, Kurumizaka H

EMDB-32409: 
RNA polymerase II elongation complex bound with Elf1 and Spt4/5, stalled at SHL(-3) of the nucleosome
Method: single particle / : Osumi K, Kujirai T, Ehara H, Sekine S, Takizawa Y, Kurumizaka H

EMDB-34685: 
RNA polymerase II elongation complex bound with Rad26 and Elf1, stalled at SHL(-3.5) of the nucleosome
Method: single particle / : Osumi K, Kujirai T, Ehara H, Kinoshita C, Saotome M, Kagawa W, Sekine S, Takizawa Y, Kurumizaka H

PDB-7wbv: 
RNA polymerase II elongation complex bound with Elf1 and Spt4/5, stalled at SHL(-4) of the nucleosome
Method: single particle / : Osumi K, Kujirai T, Ehara H, Sekine S, Takizawa Y, Kurumizaka H

PDB-7wbw: 
RNA polymerase II elongation complex bound with Elf1 and Spt4/5, stalled at SHL(-3.5) of the nucleosome
Method: single particle / : Osumi K, Kujirai T, Ehara H, Sekine S, Takizawa Y, Kurumizaka H

PDB-7wbx: 
RNA polymerase II elongation complex bound with Elf1 and Spt4/5, stalled at SHL(-3) of the nucleosome
Method: single particle / : Osumi K, Kujirai T, Ehara H, Sekine S, Takizawa Y, Kurumizaka H

PDB-8he5: 
RNA polymerase II elongation complex bound with Rad26 and Elf1, stalled at SHL(-3.5) of the nucleosome
Method: single particle / : Osumi K, Kujirai T, Ehara H, Kinoshita C, Saotome M, Kagawa W, Sekine S, Takizawa Y, Kurumizaka H
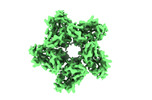
EMDB-35143: 
Cryo-EM structure of the zeaxanthin-bound kin4B8
Method: single particle / : Murakoshi S, Chazan A, Shihoya W, Beja O, Nureki O

PDB-8i2z: 
Cryo-EM structure of the zeaxanthin-bound kin4B8
Method: single particle / : Murakoshi S, Chazan A, Shihoya W, Beja O, Nureki O

EMDB-34430: 
Cryo-EM structure of the human RAD52 protein
Method: single particle / : Kinoshita C, Takizawa Y, Saotome M, Ogino S, Kurumizaka H, Kagawa W

PDB-8h1p: 
Cryo-EM structure of the human RAD52 protein
Method: single particle / : Kinoshita C, Takizawa Y, Saotome M, Ogino S, Kurumizaka H, Kagawa W

EMDB-34031: 
Cryo-EM structure of the SARS-CoV-2 spike protein (2-up RBD) bound to neutralizing antibody NIBIC-71
Method: single particle / : Otsubo R, Minamitani T, Kobiyama K, Fujita J, Ito T, Ueno S, Anzai I, Tanino H, Aoyama H, Matsuura Y, Namba K, Imadome K, Ishii KJ, Tsumoto K, Kamitani W, Yasui T

EMDB-34032: 
Cryo-EM structure of the SARS-CoV-2 spike protein (1-up RBD) bound to neutralizing antibody 7G7.2
Method: single particle / : Otsubo R, Minamitani T, Kobiyama K, Fujita J, Ito T, Ueno S, Anzai I, Tanino H, Aoyama H, Matsuura Y, Namba K, Imadome K, Ishii KJ, Tsumoto K, Kamitani W, Yasui T

EMDB-34097: 
Human Lysophosphatidic Acid Receptor 1-Gi complex bound to ONO-0740556
Method: single particle / : Akasaka H, Shihoya W, Nureki O

EMDB-34098: 
Human Lysophosphatidic Acid Receptor 1-Gi complex bound to ONO-0740556, focused on receptor
Method: single particle / : Akasaka H, Shihoya W, Nureki O

EMDB-34099: 
Human Lysophosphatidic Acid Receptor 1-Gi complex bound to ONO-0740556, state1
Method: single particle / : Akasaka H, Shihoya W, Nureki O

EMDB-34100: 
Human Lysophosphatidic Acid Receptor 1-Gi complex bound to ONO-0740556, state2
Method: single particle / : Akasaka H, Shihoya W, Nureki O

EMDB-34101: 
Human Lysophosphatidic Acid Receptor 1-Gi complex bound to ONO-0740556, state3
Method: single particle / : Akasaka H, Shihoya W, Nureki O

EMDB-34102: 
Human Lysophosphatidic Acid Receptor 1-Gi complex bound to ONO-0740556, state4
Method: single particle / : Akasaka H, Shihoya W, Nureki O
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
